



Next: List of Tables
Up: EXPERIMENTATION AT LINEAR COLLIDERS
Previous: Contents
-
Requirements of luminosity as a function of the
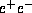 center-of-mass energy of
linear colliders, where the energies of the first and second phases and the minimum
required luminosities are depicted together with the physics targets.
center-of-mass energy of
linear colliders, where the energies of the first and second phases and the minimum
required luminosities are depicted together with the physics targets.
-
Top view of the beam line from the exit of the main linac to the
interaction point (IP) of JLC-I(
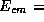 500GeV).
500GeV).
- Muon attenuator for JLC-I(KEK) presented by Y.Namito at LC93. Two iron pipes are
magnetized axially in opposite directions for both charged muons which can be trapped;
also, the 120m length of the iron pipe corresponds to a mean range of 250GeV muons.
- Muon spoiler for NLC(SLAC) presented by L.Keller at LC93. A square tunnel of
3x3m
 is filled by the 1.5 Tesla magnetized spoiler, the length and weight of which are
9.1m and 750 tons, respectively.
is filled by the 1.5 Tesla magnetized spoiler, the length and weight of which are
9.1m and 750 tons, respectively.
- Horizontal(6
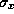 ) and vertical(35
) and vertical(35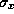 ) beam envelopes through the
last bending magnet and four final focus quadrupole magnets(QC1,QC2,QC3,QC4) for the JLC-I.
The maximum divergences of the synchrotron radiation are also drawn by arrows.
) beam envelopes through the
last bending magnet and four final focus quadrupole magnets(QC1,QC2,QC3,QC4) for the JLC-I.
The maximum divergences of the synchrotron radiation are also drawn by arrows.
- Profiles of the synchrotron radiation at QC1.
The profile at the center accompanies the in-coming beam. The two right-hand
side's ones are passing through QC1 of 2.4m long after a collision with a 8mrad
horizontal crossing.
- Electrons and positrons scattered by beam-beam interaction at JLC-I, where the
the vertical and horizontal axes are the scattering angles in vertical and horizontal
directions, respectively.
- Masking system in a solenoidal magnetic field of 2 Tesla around the IP for
JLC-I.
- Particles of
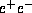 pairs at the front face of the mask (44cm from the IP)
for JLC-I, where the vertical axis is their energy and the horizontal one is their
radial position.
pairs at the front face of the mask (44cm from the IP)
for JLC-I, where the vertical axis is their energy and the horizontal one is their
radial position.
- Hits per a bunch crossing by the pairs in B=2Tesla as a function of the
radius(
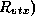 of a cylinder for two angular regions ((a) and (b)) for JLC-I.
of a cylinder for two angular regions ((a) and (b)) for JLC-I.
- Masking system for TESLA proposed by D.Schulte at LC95, where
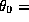 66mrad and
66mrad and 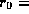 33mm. The nearest superconducting quadrupole magnet is
located 3m from the IP, and the aperture is 48mm
33mm. The nearest superconducting quadrupole magnet is
located 3m from the IP, and the aperture is 48mm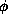 .
.
- Hits per a bunch crossing in the inner layer of the vertex detector for TESLA,
where the vertex detector covers the angular region
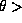 200mrad.
200mrad.
- Schematic view of the JLC detector. The total volume and weight are
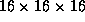 m
m and 15,000 ton, respectively.
and 15,000 ton, respectively.
- Power-spectrum density of vertical ground motion at various locations.
- Ground-motion tolerances and
integrated spectrum as a function of the frequency, based on SLC linac measurements during
``quiet" conditions.
Toshiaki Tauchi
Fri, Dec 20, 1996 02:24:05 PM